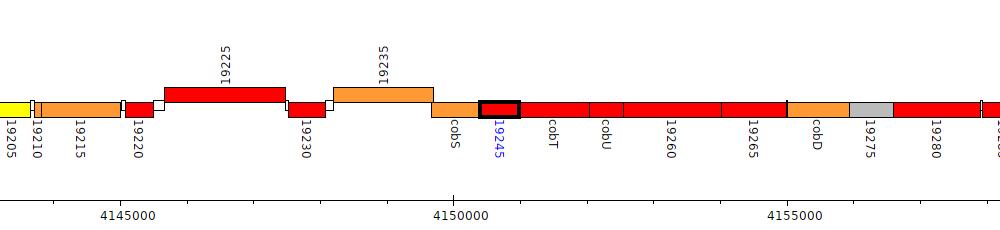
Pseudomonas aeruginosa DK2, PADK2_19245
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pdk00860 | Porphyrin and chlorophyll metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pdk01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR48100 | BROAD-SPECIFICITY PHOSPHATASE YOR283W-RELATED | - | - | 4 | 164 | 5.8E-19 |
| SUPERFAMILY | SSF53254 | Phosphoglycerate mutase-like | IPR029033 | Histidine phosphatase superfamily | 2 | 188 | 7.07E-48 |
| PIRSF | PIRSF000709 | 6PFK_fruc_bisph_Ptase | - | - | 6 | 189 | 0.0056 |
| Gene3D | G3DSA:3.40.50.1240 | - | IPR029033 | Histidine phosphatase superfamily | 2 | 191 | 5.0E-46 |
| CDD | cd07067 | HP_PGM_like | IPR013078 | Histidine phosphatase superfamily, clade-1 | 4 | 175 | 1.31236E-23 |
| SMART | SM00855 | PGAM_5 | IPR013078 | Histidine phosphatase superfamily, clade-1 | 4 | 153 | 5.6E-15 |
| Pfam | PF00300 | Histidine phosphatase superfamily (branch 1) | IPR013078 | Histidine phosphatase superfamily, clade-1 | 7 | 184 | 1.5E-42 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.