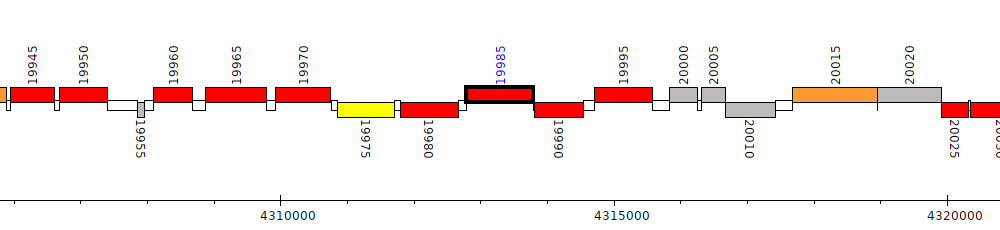
Pseudomonas aeruginosa DK2, PADK2_19985
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 130 | 273 | 9.5E-107 |
| Pfam | PF13602 | Zinc-binding dehydrogenase | - | - | 194 | 334 | 7.0E-26 |
| SMART | SM00829 | PKS_ER_names_mod | IPR020843 | Polyketide synthase, enoylreductase domain | 20 | 334 | 2.8E-59 |
| CDD | cd05276 | p53_inducible_oxidoreductase | IPR014189 | Quinone oxidoreductase PIG3 | 11 | 334 | 0.0 |
| Pfam | PF08240 | Alcohol dehydrogenase GroES-like domain | IPR013154 | Alcohol dehydrogenase-like, N-terminal | 38 | 97 | 2.4E-9 |
| PANTHER | PTHR48106 | QUINONE OXIDOREDUCTASE PIG3-RELATED | - | - | 11 | 335 | 2.7E-93 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 121 | 298 | 1.85E-47 |
| NCBIfam | TIGR02824 | JCVI: putative NAD(P)H quinone oxidoreductase, PIG3 family | IPR014189 | Quinone oxidoreductase PIG3 | 11 | 335 | 0.0 |
| SUPERFAMILY | SSF50129 | GroES-like | IPR011032 | GroES-like superfamily | 11 | 150 | 7.15E-39 |
| Gene3D | G3DSA:3.90.180.10 | - | - | - | 22 | 334 | 9.5E-107 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.