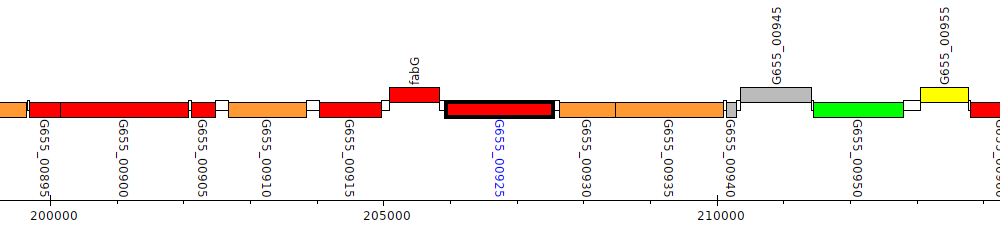
Pseudomonas aeruginosa B136-33, G655_00925
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004065 | arylsulfatase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0183
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
7744061 | |
| Molecular Function | GO:0008081 | phosphoric diester hydrolase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0183
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
19554727 | |
| Molecular Function | GO:0008484 | sulfuric ester hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psg00600 | Sphingolipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.1120.10 | - | - | - | 451 | 536 | 8.5E-37 |
| CDD | cd16025 | PAS_like | - | - | 3 | 502 | 0.0 |
| PANTHER | PTHR42693 | ARYLSULFATASE FAMILY MEMBER | - | - | 2 | 523 | 1.3E-107 |
| Gene3D | G3DSA:3.40.720.10 | Alkaline Phosphatase, subunit A | IPR017850 | Alkaline-phosphatase-like, core domain superfamily | 1 | 450 | 0.0 |
| FunFam | G3DSA:3.40.720.10:FF:000044 | Arylsulfatase | - | - | 1 | 449 | 0.0 |
| Pfam | PF00884 | Sulfatase | IPR000917 | Sulfatase, N-terminal | 5 | 420 | 3.1E-91 |
| SUPERFAMILY | SSF53649 | Alkaline phosphatase-like | IPR017850 | Alkaline-phosphatase-like, core domain superfamily | 3 | 526 | 0.0 |
| FunFam | G3DSA:3.30.1120.10:FF:000005 | Arylsulfatase A | - | - | 451 | 533 | 6.3E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.