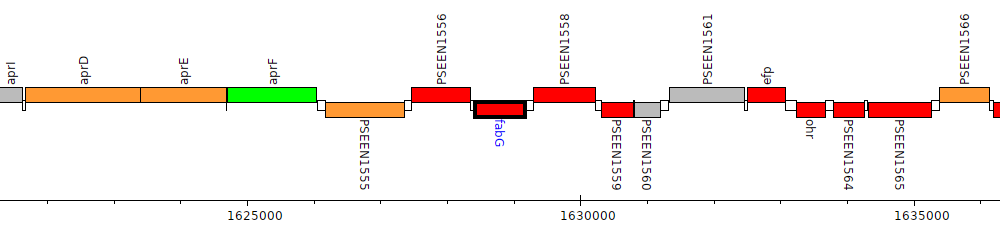
Pseudomonas entomophila L48, PSEEN1557 (fabG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pen01212 | Fatty acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen01040 | Biosynthesis of unsaturated fatty acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00061 | Fatty acid biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00780 | Biotin metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 131 | 147 | 3.1E-40 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 2 | 248 | 4.4E-82 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 158 | 177 | 2.4E-13 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 211 | 231 | 3.1E-40 |
| CDD | cd05233 | SDR_c | - | - | 12 | 244 | 1.25246E-76 |
| Pfam | PF13561 | Enoyl-(Acyl carrier protein) reductase | - | - | 18 | 247 | 5.2E-62 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 86 | 97 | 2.4E-13 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 158 | 177 | 3.1E-40 |
| SMART | SM00822 | This enzymatic domain is part of bacterial polyketide synthases and catalyses the first step in the reductive modification of the beta-carbonyl centres in the growing polyketide chain. It uses NADPH to reduce the keto group to a hydroxy group. | - | - | 10 | 189 | 0.0018 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 11 | 28 | 3.1E-40 |
| FunFam | G3DSA:3.40.50.720:FF:000290 | SDR family oxidoreductase | - | - | 1 | 248 | 3.4E-111 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 86 | 97 | 3.1E-40 |
| PANTHER | PTHR43639 | OXIDOREDUCTASE, SHORT-CHAIN DEHYDROGENASE/REDUCTASE FAMILY (AFU_ORTHOLOGUE AFUA_5G02870) | - | - | 2 | 246 | 1.3E-67 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 5 | 248 | 6.16E-80 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 137 | 145 | 2.4E-13 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 179 | 196 | 3.1E-40 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.