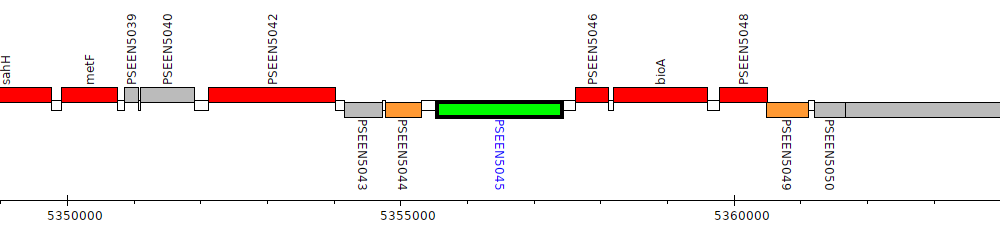
Pseudomonas entomophila L48, PSEEN5045
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00757
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pen00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00380 | Tryptophan metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00340 | Histidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00350 | Tyrosine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00360 | Phenylalanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00757 | Flavin-containing amine oxidase signature | IPR001613 | Flavin amine oxidase | 432 | 449 | 1.8E-10 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 24 | 452 | 1.78E-50 |
| SUPERFAMILY | SSF54373 | FAD-linked reductases, C-terminal domain | - | - | 295 | 399 | 2.12E-14 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 560 | 620 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 592 | 606 | - |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 22 | 458 | 2.3E-80 |
| PRINTS | PR00757 | Flavin-containing amine oxidase signature | IPR001613 | Flavin amine oxidase | 401 | 423 | 1.8E-10 |
| Pfam | PF01593 | Flavin containing amine oxidoreductase | IPR002937 | Amine oxidase | 35 | 450 | 8.6E-45 |
| PANTHER | PTHR10742 | FLAVIN MONOAMINE OXIDASE | - | - | 28 | 453 | 6.1E-38 |
| PRINTS | PR00757 | Flavin-containing amine oxidase signature | IPR001613 | Flavin amine oxidase | 27 | 46 | 1.8E-10 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.