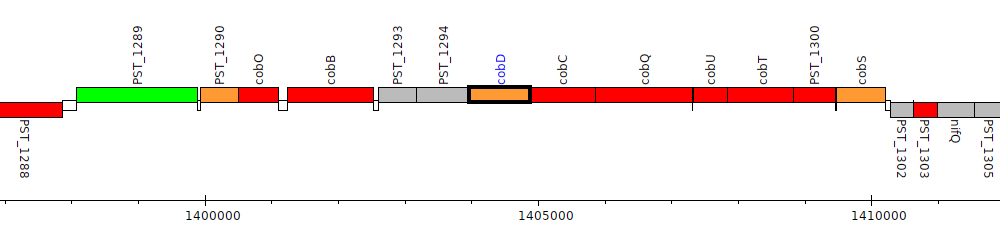
Pseudomonas stutzeri A1501, PST_1295 (cobD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009236 | cobalamin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF03186
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF03186
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0048472 | threonine-phosphate decarboxylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF03186
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psa01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa00860 | Porphyrin and chlorophyll metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00380 | JCVI: adenosylcobinamide-phosphate synthase CbiB | IPR004485 | Cobalamin biosynthesis CobD/CbiB | 7 | 293 | 5.2E-61 |
| Pfam | PF17113 | Regulatory signalling modulator protein AmpE | IPR031347 | Protein AmpE | 150 | 225 | 7.8E-7 |
| PANTHER | PTHR34308 | COBALAMIN BIOSYNTHESIS PROTEIN CBIB | IPR004485 | Cobalamin biosynthesis CobD/CbiB | 4 | 296 | 1.2E-101 |
| Hamap | MF_00024 | Cobalamin biosynthesis protein CbiB [cbiB]. | IPR004485 | Cobalamin biosynthesis CobD/CbiB | 1 | 303 | 108.063766 |
| Pfam | PF03186 | CobD/Cbib protein | IPR004485 | Cobalamin biosynthesis CobD/CbiB | 8 | 284 | 1.2E-95 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.