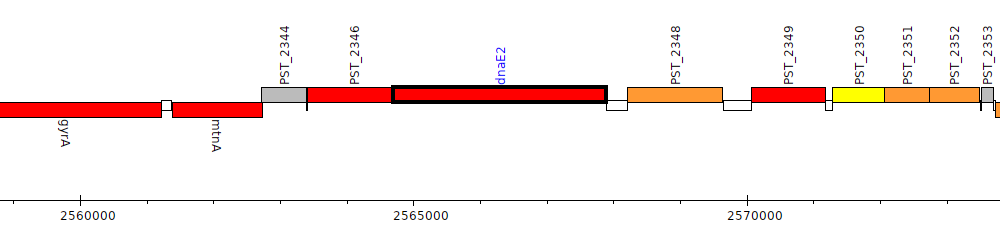
Pseudomonas stutzeri A1501, PST_2347 (dnaE2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02811
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32294
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008408 | 3'-5' exonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32294
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01902
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.150.870 | - | - | - | 795 | 884 | 4.5E-11 |
| CDD | cd04485 | DnaE_OBF | - | - | 977 | 1063 | 2.36226E-18 |
| NCBIfam | TIGR00594 | JCVI: DNA polymerase III subunit alpha | IPR004805 | Error-prone DNA polymerase/DNA polymerase III subunit alpha DnaE/PolC | 45 | 1010 | 0.0 |
| Hamap | MF_01902 | Error-prone DNA polymerase [dnaE2]. | IPR023073 | DNA polymerase, error-prone | 44 | 1065 | 13.752165 |
| Pfam | PF07733 | Bacterial DNA polymerase III alpha NTPase domain | IPR011708 | Bacterial DNA polymerase III, alpha subunit, NTPase domain | 310 | 564 | 6.5E-83 |
| FunFam | G3DSA:1.10.150.870:FF:000002 | Error-prone DNA polymerase | - | - | 792 | 884 | 4.5E-32 |
| Pfam | PF17657 | Bacterial DNA polymerase III alpha subunit finger domain | IPR040982 | DNA polymerase III alpha subunit finger domain | 567 | 732 | 1.3E-59 |
| SUPERFAMILY | SSF89550 | PHP domain-like | IPR016195 | Polymerase/histidinol phosphatase-like | 44 | 227 | 5.49E-14 |
| CDD | cd07434 | PHP_PolIIIA_DnaE2 | - | - | 44 | 297 | 2.31227E-124 |
| Pfam | PF02811 | PHP domain | IPR004013 | PHP domain | 46 | 163 | 1.2E-15 |
| Pfam | PF01336 | OB-fold nucleic acid binding domain | IPR004365 | OB-fold nucleic acid binding domain, AA-tRNA synthetase-type | 975 | 1042 | 3.2E-8 |
| Gene3D | G3DSA:3.20.20.140 | - | - | - | 25 | 287 | 6.2E-59 |
| SMART | SM00481 | npolultra | IPR003141 | Polymerase/histidinol phosphatase, N-terminal | 45 | 112 | 4.4E-10 |
| PANTHER | PTHR32294 | DNA POLYMERASE III SUBUNIT ALPHA | IPR004805 | Error-prone DNA polymerase/DNA polymerase III subunit alpha DnaE/PolC | 40 | 1053 | 0.0 |
| Pfam | PF14579 | Helix-hairpin-helix motif | IPR029460 | DNA polymerase, helix-hairpin-helix motif | 807 | 895 | 1.8E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.