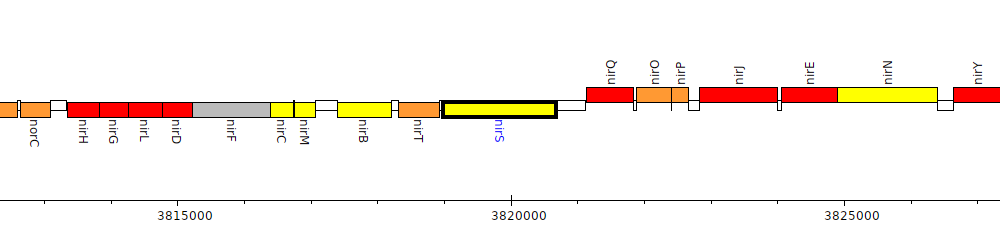
Pseudomonas stutzeri A1501, PST_3532 (nirS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF13442
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF13442
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psa00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psa01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.760.10 | - | IPR036909 | Cytochrome c-like domain superfamily | 21 | 127 | 6.8E-23 |
| SUPERFAMILY | SSF51004 | C-terminal (heme d1) domain of cytochrome cd1-nitrite reductase | IPR011048 | Cytochrome cd1-nitrite reductase-like, haem d1 domain superfamily | 129 | 560 | 9.29E-121 |
| Pfam | PF13442 | Cytochrome C oxidase, cbb3-type, subunit III | IPR009056 | Cytochrome c-like domain | 37 | 120 | 4.2E-9 |
| Pfam | PF02239 | Cytochrome D1 heme domain | - | - | 157 | 555 | 0.0 |
| Gene3D | G3DSA:2.140.10.20 | - | IPR003143 | Cytochrome cd1-nitrite reductase, C-terminal domain superfamily | 129 | 560 | 0.0 |
| SUPERFAMILY | SSF46626 | Cytochrome c | IPR036909 | Cytochrome c-like domain superfamily | 12 | 141 | 7.74E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.