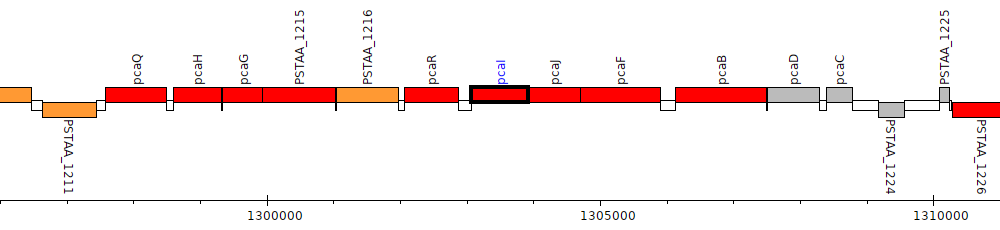
Pseudomonas stutzeri DSM 4166, PSTAA_1218 (pcaI)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008410 | CoA-transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01144
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psr00643 | Styrene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psr00650 | Butanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psr01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.1080.10 | - | - | - | 5 | 281 | 2.6E-71 |
| SUPERFAMILY | SSF100950 | NagB/RpiA/CoA transferase-like | IPR037171 | NagB/RpiA transferase-like | 5 | 278 | 1.41E-79 |
| SMART | SM00882 | CoA_trans_3 | IPR004165 | Coenzyme A transferase family I | 4 | 224 | 1.3E-43 |
| Coils | Coil | Coil | - | - | 271 | 285 | - |
| Pfam | PF01144 | Coenzyme A transferase | IPR004165 | Coenzyme A transferase family I | 6 | 215 | 9.2E-25 |
| Gene3D | G3DSA:3.30.30.40 | - | - | - | 118 | 154 | 2.6E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.