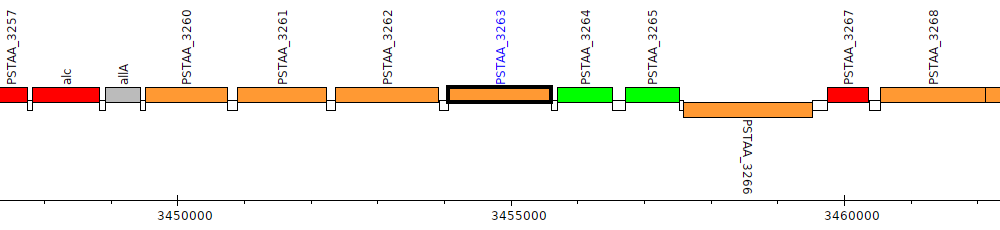
Pseudomonas stutzeri DSM 4166, PSTAA_3263
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00801
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00801
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015851 | nucleobase transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00801
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | NF037981 | NCBIFAM: purine/pyrimidine permease | - | - | 45 | 456 | 9.2E-14 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 491 | 513 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 22 | - |
| PANTHER | PTHR42810 | PURINE PERMEASE C1399.01C-RELATED | - | - | 25 | 468 | 0.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 491 | 505 | - |
| NCBIfam | TIGR03173 | JCVI: xanthine permease | IPR017588 | Uric acid permease UacT-like | 46 | 461 | 9.9E-129 |
| Pfam | PF00860 | Permease family | IPR006043 | Nucleobase cation symporter 2 family | 43 | 429 | 1.4E-83 |
| NCBIfam | TIGR00801 | JCVI: NCS2 family nucleobase:cation symporter | IPR006042 | Xanthine/uracil permease | 40 | 458 | 1.4E-111 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.