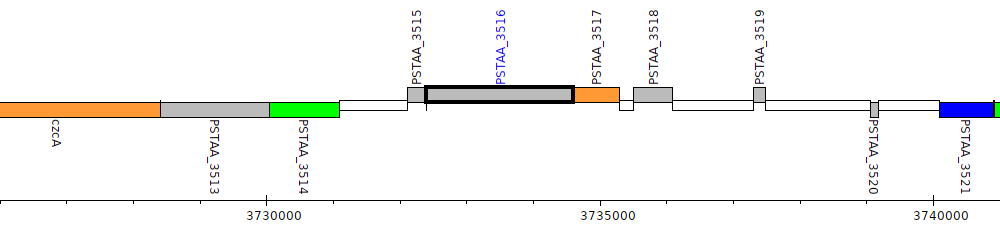
Pseudomonas stutzeri DSM 4166, PSTAA_3516
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00175
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00111
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06217 | FNR_iron_sulfur_binding_3 | - | - | 389 | 624 | 1.10319E-107 |
| PRINTS | PR00371 | Flavoprotein pyridine nucleotide cytochrome reductase signature | IPR001709 | Flavoprotein pyridine nucleotide cytochrome reductase | 538 | 549 | 9.3E-7 |
| SUPERFAMILY | SSF52343 | Ferredoxin reductase-like, C-terminal NADP-linked domain | IPR039261 | Ferredoxin-NADP reductase (FNR), nucleotide-binding domain | 481 | 622 | 6.29E-37 |
| PRINTS | PR00410 | Phenol hydroxylase reductase family signature | - | - | 501 | 520 | 6.7E-10 |
| CDD | cd00207 | fer2 | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 646 | 729 | 3.06527E-22 |
| Pfam | PF00111 | 2Fe-2S iron-sulfur cluster binding domain | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 654 | 722 | 5.3E-13 |
| Gene3D | G3DSA:3.40.50.80 | - | IPR039261 | Ferredoxin-NADP reductase (FNR), nucleotide-binding domain | 493 | 627 | 6.2E-32 |
| SUPERFAMILY | SSF54292 | 2Fe-2S ferredoxin-like | IPR036010 | 2Fe-2S ferredoxin-like superfamily | 618 | 729 | 6.95E-25 |
| SUPERFAMILY | SSF63380 | Riboflavin synthase domain-like | IPR017938 | Riboflavin synthase-like beta-barrel | 391 | 493 | 1.44E-25 |
| PRINTS | PR00410 | Phenol hydroxylase reductase family signature | - | - | 592 | 600 | 6.7E-10 |
| PRINTS | PR00410 | Phenol hydroxylase reductase family signature | - | - | 422 | 434 | 6.7E-10 |
| PRINTS | PR00371 | Flavoprotein pyridine nucleotide cytochrome reductase signature | IPR001709 | Flavoprotein pyridine nucleotide cytochrome reductase | 441 | 448 | 9.3E-7 |
| PRINTS | PR00371 | Flavoprotein pyridine nucleotide cytochrome reductase signature | IPR001709 | Flavoprotein pyridine nucleotide cytochrome reductase | 501 | 520 | 9.3E-7 |
| Pfam | PF00970 | Oxidoreductase FAD-binding domain | IPR008333 | Flavoprotein pyridine nucleotide cytochrome reductase-like, FAD-binding domain | 420 | 491 | 6.7E-13 |
| Gene3D | G3DSA:3.10.20.30 | - | IPR012675 | Beta-grasp domain superfamily | 636 | 730 | 2.6E-22 |
| PANTHER | PTHR47354 | NADH OXIDOREDUCTASE HCR | - | - | 299 | 729 | 2.5E-96 |
| Pfam | PF00175 | Oxidoreductase NAD-binding domain | IPR001433 | Oxidoreductase FAD/NAD(P)-binding | 502 | 606 | 2.7E-17 |
| PRINTS | PR00410 | Phenol hydroxylase reductase family signature | - | - | 485 | 494 | 6.7E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 107 | 121 | - |
| PRINTS | PR00371 | Flavoprotein pyridine nucleotide cytochrome reductase signature | IPR001709 | Flavoprotein pyridine nucleotide cytochrome reductase | 526 | 535 | 9.3E-7 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 80 | 160 | - |
| Gene3D | G3DSA:2.40.30.10 | Translation factors | - | - | 389 | 492 | 2.7E-24 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.