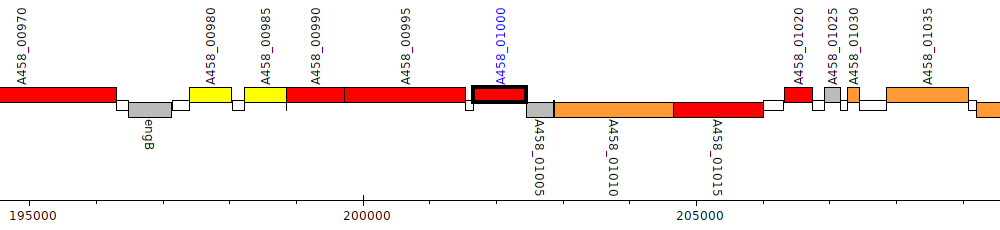
Pseudomonas stutzeri CCUG 29243, A458_01000
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009253 | peptidoglycan catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01510
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01510
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF47090 | PGBD-like | IPR036365 | PGBD-like superfamily | 186 | 256 | 2.35E-18 |
| SUPERFAMILY | SSF55846 | N-acetylmuramoyl-L-alanine amidase-like | IPR036505 | N-acetylmuramoyl-L-alanine amidase/PGRP domain superfamily | 21 | 175 | 1.83E-49 |
| FunFam | G3DSA:3.40.80.10:FF:000003 | N-acetylmuramoyl-L-alanine amidase | - | - | 33 | 178 | 1.4E-58 |
| Gene3D | G3DSA:3.40.80.10 | - | IPR036505 | N-acetylmuramoyl-L-alanine amidase/PGRP domain superfamily | 33 | 178 | 3.5E-50 |
| Gene3D | G3DSA:1.10.101.10 | - | IPR036366 | PGBD superfamily | 179 | 258 | 1.4E-26 |
| Pfam | PF01471 | Putative peptidoglycan binding domain | IPR002477 | Peptidoglycan binding-like | 202 | 253 | 8.7E-7 |
| Pfam | PF01510 | N-acetylmuramoyl-L-alanine amidase | IPR002502 | N-acetylmuramoyl-L-alanine amidase domain | 34 | 165 | 3.5E-27 |
| PANTHER | PTHR30417 | N-ACETYLMURAMOYL-L-ALANINE AMIDASE AMID | - | - | 25 | 255 | 2.0E-56 |
| SMART | SM00644 | ami_2 | IPR002502 | N-acetylmuramoyl-L-alanine amidase domain | 23 | 164 | 1.7E-30 |
| CDD | cd06583 | PGRP | IPR002502 | N-acetylmuramoyl-L-alanine amidase domain | 34 | 166 | 1.80216E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.