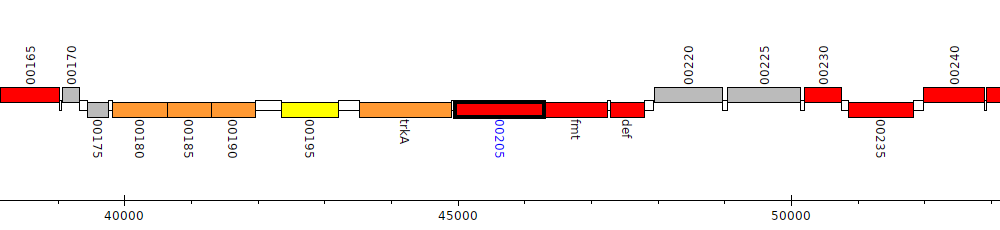
Pseudomonas stutzeri DSM 10701, PSJM300_00205
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF01029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01189
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0001510 | RNA methylation |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR22807
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008649 | rRNA methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006364 | rRNA processing |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR22807 | NOP2 YEAST -RELATED NOL1/NOP2/FMU SUN DOMAIN-CONTAINING | IPR023267 | RNA (C5-cytosine) methyltransferase | 126 | 435 | 4.1E-62 |
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 231 | 434 | 5.5E-65 |
| Coils | Coil | Coil | - | - | 284 | 304 | - |
| PRINTS | PR02008 | RNA (C5-cytosine) methyltransferase signature | IPR023267 | RNA (C5-cytosine) methyltransferase | 318 | 330 | 2.4E-28 |
| SUPERFAMILY | SSF48013 | NusB-like | IPR035926 | NusB-like superfamily | 4 | 149 | 1.06E-32 |
| FunFam | G3DSA:3.40.50.150:FF:000022 | Ribosomal RNA small subunit methyltransferase B | - | - | 234 | 434 | 2.4E-73 |
| PRINTS | PR02008 | RNA (C5-cytosine) methyltransferase signature | IPR023267 | RNA (C5-cytosine) methyltransferase | 222 | 236 | 2.4E-28 |
| Pfam | PF01029 | NusB family | IPR006027 | NusB/RsmB/TIM44 | 4 | 126 | 7.8E-19 |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 138 | 434 | 6.54E-73 |
| PRINTS | PR02008 | RNA (C5-cytosine) methyltransferase signature | IPR023267 | RNA (C5-cytosine) methyltransferase | 416 | 433 | 2.4E-28 |
| NCBIfam | TIGR00563 | JCVI: 16S rRNA (cytosine(967)-C(5))-methyltransferase RsmB | IPR004573 | rRNA small subunit methyltransferase B | 10 | 433 | 2.2E-124 |
| Gene3D | G3DSA:1.10.940.10 | - | IPR035926 | NusB-like superfamily | 3 | 143 | 2.5E-31 |
| PRINTS | PR02008 | RNA (C5-cytosine) methyltransferase signature | IPR023267 | RNA (C5-cytosine) methyltransferase | 251 | 261 | 2.4E-28 |
| CDD | cd02440 | AdoMet_MTases | - | - | 249 | 373 | 5.06002E-7 |
| PRINTS | PR02008 | RNA (C5-cytosine) methyltransferase signature | IPR023267 | RNA (C5-cytosine) methyltransferase | 368 | 384 | 2.4E-28 |
| Gene3D | G3DSA:1.10.287.730 | Helix hairpin bin | - | - | 146 | 172 | 1.0E-9 |
| Gene3D | G3DSA:3.30.70.1170 | Sun protein; domain 3 | - | - | 173 | 230 | 5.3E-23 |
| FunFam | G3DSA:3.30.70.1170:FF:000002 | Ribosomal RNA small subunit methyltransferase B | - | - | 173 | 230 | 7.8E-21 |
| Pfam | PF01189 | 16S rRNA methyltransferase RsmB/F | IPR001678 | SAM-dependent methyltransferase RsmB/NOP2-type | 240 | 431 | 1.3E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.