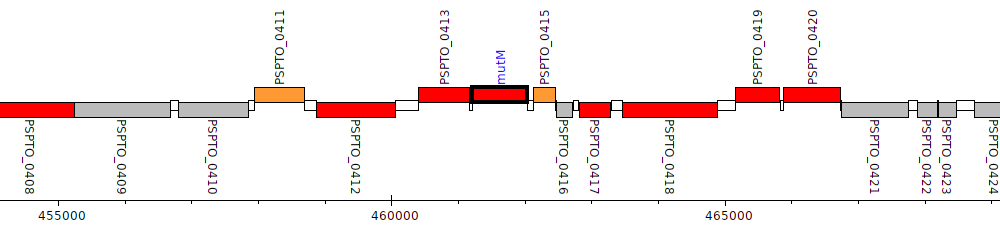
Pseudomonas syringae pv. tomato DC3000, PSPTO_0414 (mutM)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0019104 | DNA N-glycosylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46946
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003906 | DNA-(apurinic or apyrimidinic site) endonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00577
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003684 | damaged DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM01232
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006284 | base-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:SM00898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008534 | oxidized purine nucleobase lesion DNA N-glycosylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00577
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00577
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016799 | hydrolase activity, hydrolyzing N-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:SM01232
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00577
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pst03410 | Base excision repair | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd08966 | EcFpg-like_N | - | - | 2 | 117 | 1.65483E-50 |
| SUPERFAMILY | SSF81624 | N-terminal domain of MutM-like DNA repair proteins | IPR035937 | MutM-like, N-terminal | 2 | 135 | 7.46E-40 |
| FunFam | G3DSA:1.10.8.50:FF:000003 | Formamidopyrimidine-DNA glycosylase | - | - | 132 | 270 | 2.6E-49 |
| PANTHER | PTHR22993 | FORMAMIDOPYRIMIDINE-DNA GLYCOSYLASE | - | - | 1 | 270 | 1.3E-44 |
| FunFam | G3DSA:3.20.190.10:FF:000001 | Formamidopyrimidine-DNA glycosylase | - | - | 2 | 129 | 9.5E-47 |
| Hamap | MF_00103 | Formamidopyrimidine-DNA glycosylase [mutM]. | IPR020629 | Formamidopyrimidine-DNA glycosylase | 1 | 270 | 44.101097 |
| SMART | SM01232 | H2TH_2 | IPR015886 | DNA glycosylase/AP lyase, H2TH DNA-binding | 130 | 222 | 1.9E-42 |
| Pfam | PF06831 | Formamidopyrimidine-DNA glycosylase H2TH domain | IPR015886 | DNA glycosylase/AP lyase, H2TH DNA-binding | 130 | 220 | 2.3E-24 |
| SMART | SM00898 | Fapy_DNA_glyco_2 | IPR012319 | Formamidopyrimidine-DNA glycosylase, catalytic domain | 2 | 116 | 1.9E-52 |
| Gene3D | G3DSA:1.10.8.50 | - | - | - | 132 | 270 | 3.1E-43 |
| Pfam | PF01149 | Formamidopyrimidine-DNA glycosylase N-terminal domain | IPR012319 | Formamidopyrimidine-DNA glycosylase, catalytic domain | 1 | 115 | 5.2E-33 |
| SUPERFAMILY | SSF46946 | S13-like H2TH domain | IPR010979 | Small ribosomal subunit protein uS13-like, H2TH | 131 | 222 | 2.22E-26 |
| NCBIfam | TIGR00577 | JCVI: DNA-formamidopyrimidine glycosylase | IPR020629 | Formamidopyrimidine-DNA glycosylase | 1 | 269 | 3.3E-90 |
| SUPERFAMILY | SSF57716 | Glucocorticoid receptor-like (DNA-binding domain) | - | - | 218 | 269 | 2.51E-17 |
| Pfam | PF06827 | Zinc finger found in FPG and IleRS | IPR010663 | Zinc finger, FPG/IleRS-type | 242 | 269 | 4.1E-9 |
| Gene3D | G3DSA:3.20.190.10 | - | IPR035937 | MutM-like, N-terminal | 2 | 129 | 5.5E-40 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.