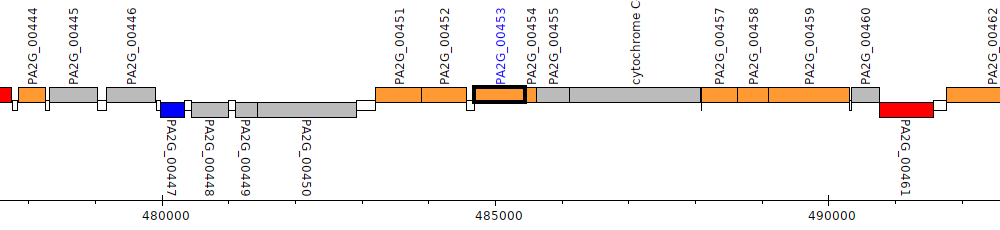
Pseudomonas aeruginosa 2192, PA2G_00453
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071978 | bacterial-type flagellum-dependent swarming motility |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Biological Process | GO:0071977 | bacterial-type flagellum-dependent swimming motility |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Biological Process | GO:0002049 | pyoverdine biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Molecular Function | GO:0004096 | catalase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Biological Process | GO:0009247 | glycolipid biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Biological Process | GO:0043107 | type IV pilus-dependent motility |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1477
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
16824107 | |
| Biological Process | GO:0015886 | heme transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01191
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01191
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015232 | heme transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01191
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0017004 | cytochrome complex assembly |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR30071
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR30071
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 123 | 143 | 4.7E-59 |
| PANTHER | PTHR30071 | HEME EXPORTER PROTEIN C | IPR045062 | Cytochrome c-type biogenesis protein CcsA/CcmC | 32 | 214 | 2.2E-31 |
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 206 | 230 | 4.7E-59 |
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 47 | 64 | 4.7E-59 |
| Pfam | PF01578 | Cytochrome C assembly protein | IPR002541 | Cytochrome c assembly protein | 15 | 186 | 6.7E-26 |
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 102 | 122 | 4.7E-59 |
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 158 | 184 | 4.7E-59 |
| PRINTS | PR01386 | Cytochrome c-type biogenesis protein CcmC signature | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 64 | 84 | 4.7E-59 |
| NCBIfam | TIGR01191 | JCVI: heme ABC transporter permease CcmC | IPR003557 | Cytochrome c-type biogenesis protein CcmC | 47 | 230 | 7.1E-84 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.