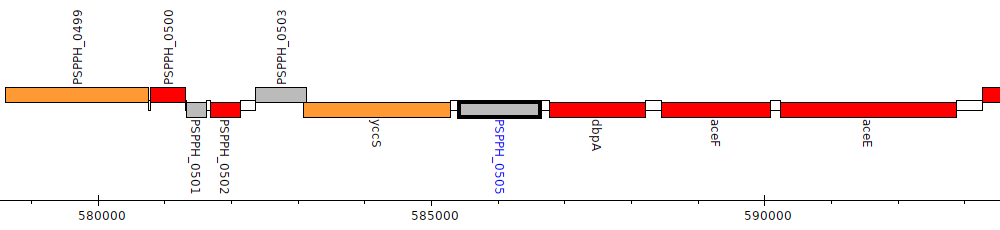
Pseudomonas savastanoi pv. phaseolicola 1448A, PSPPH_0505
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 17 | 405 | 1.73E-58 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 18 | 402 | 0.0 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 19 | 38 | 6.5E-6 |
| NCBIfam | TIGR00275 | JCVI: aminoacetone oxidase family FAD-binding enzyme | IPR004792 | 3-Dehydro-bile acid delta(4,6)-reductase-like | 19 | 403 | 9.2E-128 |
| Gene3D | G3DSA:2.40.30.10 | Translation factors | - | - | 201 | 350 | 0.0 |
| PANTHER | PTHR42887 | OS12G0638800 PROTEIN | IPR004792 | 3-Dehydro-bile acid delta(4,6)-reductase-like | 17 | 404 | 1.4E-93 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 168 | 177 | 6.2E-9 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 18 | 40 | 6.2E-9 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 358 | 380 | 6.5E-6 |
| Pfam | PF03486 | HI0933-like protein | - | - | 18 | 403 | 0.0 |
| SUPERFAMILY | SSF160996 | HI0933 insert domain-like | - | - | 203 | 349 | 5.49E-38 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 373 | 380 | 6.2E-9 |
| Gene3D | G3DSA:1.10.8.260 | - | IPR023166 | HI0933-like insert domain superfamily | 274 | 333 | 0.0 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 165 | 183 | 6.5E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.