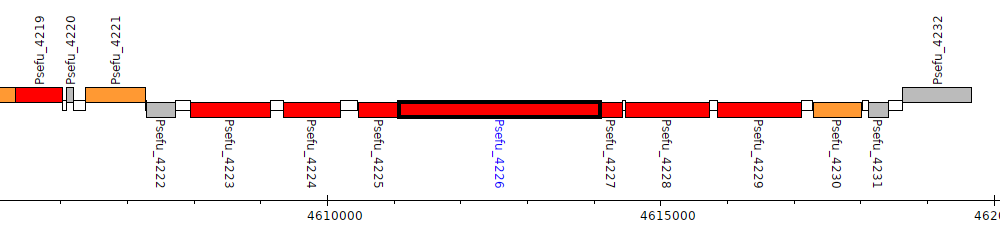
Pseudomonas fulva 12-X, Psefu_4226
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046653 | tetrahydrofolate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF037980
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1360.120
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008115 | sarcosine oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF037980
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | glycine betaine degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pfv01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfv00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | creatinine degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF101790 | Aminomethyltransferase beta-barrel domain | IPR029043 | Glycine cleavage T-protein/YgfZ, C-terminal | 907 | 1003 | 1.28E-17 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 373 | 389 | 2.9E-12 |
| Gene3D | G3DSA:3.30.1360.120 | Probable tRNA modification gtpase trme; domain 1 | IPR027266 | GTP-binding protein TrmE/Aminomethyltransferase GcvT, domain 1 | 615 | 1005 | 1.1E-129 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 295 | 423 | 2.7E-22 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 172 | 484 | 2.7E-22 |
| Gene3D | G3DSA:1.10.10.1100 | - | IPR041854 | BFD-like [2Fe-2S]-binding domain superfamily | 522 | 586 | 2.2E-5 |
| Gene3D | G3DSA:3.10.20.440 | - | IPR042204 | 2Fe-2S iron-sulfur cluster binding domain, N-terminal | 9 | 103 | 2.3E-21 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 172 | 435 | 7.4E-14 |
| PIRSF | PIRSF037980 | Sarcos_oxid_alpha | IPR006277 | Sarcosine oxidase, alpha subunit | 1 | 1005 | 0.0 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 283 | 301 | 1.8E-9 |
| SUPERFAMILY | SSF103025 | Folate-binding domain | - | - | 614 | 896 | 4.08E-92 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 284 | 292 | 2.9E-12 |
| Pfam | PF01571 | Aminomethyltransferase folate-binding domain | IPR006222 | Aminomethyltransferase, folate-binding domain | 620 | 885 | 8.4E-75 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 450 | 472 | 1.8E-9 |
| Pfam | PF13510 | 2Fe-2S iron-sulfur cluster binding domain | - | - | 17 | 102 | 9.2E-22 |
| PANTHER | PTHR43757 | AMINOMETHYLTRANSFERASE | IPR028896 | Aminomethyltransferase-like | 607 | 996 | 1.7E-56 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 167 | 470 | 1.3E-35 |
| NCBIfam | TIGR01372 | JCVI: sarcosine oxidase subunit alpha family protein | IPR006277 | Sarcosine oxidase, alpha subunit | 2 | 1005 | 0.0 |
| Pfam | PF17806 | Sarcosine oxidase A3 domain | IPR041117 | SoxA, A3 domain | 522 | 605 | 1.2E-30 |
| Pfam | PF08669 | Glycine cleavage T-protein C-terminal barrel domain | IPR013977 | Glycine cleavage T-protein, C-terminal barrel domain | 911 | 997 | 1.9E-19 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 172 | 194 | 2.9E-12 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 173 | 192 | 1.8E-9 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 460 | 478 | 2.9E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.