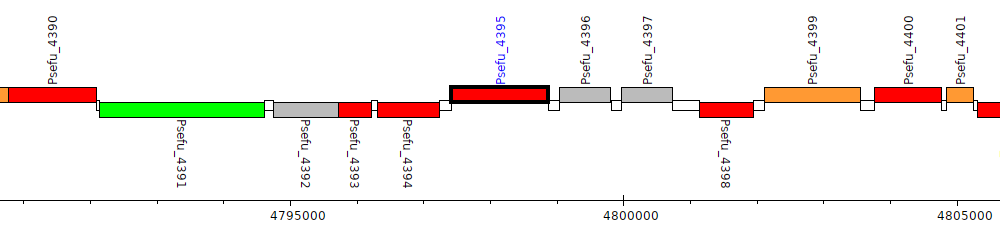
Pseudomonas fulva 12-X, Psefu_4395
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016829 | lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF10415
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:PF10415
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006531 | aspartate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00839
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008797 | aspartate ammonia-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00839
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48557
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | superpathway of glyoxylate cycle and fatty acid degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | TCA cycle V (2-oxoglutarate:ferredoxin oxidoreductase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | TCA cycle VII (acetate-producers) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | methylaspartate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | TCA cycle II (plants and fungi) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pfv01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | reductive TCA cycle II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pfv00250 | Alanine, aspartate and glutamate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | anaerobic energy metabolism (invertebrates, mitochondrial) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.200.10 | Fumarase/aspartase (Central domain) | - | - | 144 | 412 | 6.6E-81 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 322 | 338 | 1.4E-28 |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 179 | 199 | 6.6E-5 |
| Gene3D | G3DSA:1.10.40.30 | - | - | - | 413 | 470 | 1.2E-16 |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 322 | 338 | 6.6E-5 |
| FunFam | G3DSA:1.10.40.30:FF:000002 | Fumarate hydratase class II | - | - | 413 | 469 | 1.6E-16 |
| CDD | cd01357 | Aspartase | - | - | 8 | 459 | 0.0 |
| SUPERFAMILY | SSF48557 | L-aspartase-like | IPR008948 | L-Aspartase-like | 7 | 465 | 0.0 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 184 | 202 | 1.4E-28 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 276 | 303 | 1.4E-28 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 139 | 157 | 1.4E-28 |
| PANTHER | PTHR42696 | ASPARTATE AMMONIA-LYASE | - | - | 6 | 469 | 0.0 |
| FunFam | G3DSA:1.20.200.10:FF:000001 | Fumarate hydratase, mitochondrial | - | - | 144 | 411 | 4.2E-113 |
| FunFam | G3DSA:1.10.275.10:FF:000001 | Fumarate hydratase, mitochondrial | - | - | 8 | 143 | 2.1E-52 |
| Gene3D | G3DSA:1.10.275.10 | - | IPR024083 | Fumarase/histidase, N-terminal | 5 | 143 | 2.3E-53 |
| Pfam | PF00206 | Lyase | IPR022761 | Fumarate lyase, N-terminal | 15 | 347 | 3.8E-99 |
| Pfam | PF10415 | Fumarase C C-terminus | IPR018951 | Fumarase C, C-terminal | 413 | 465 | 1.1E-19 |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 138 | 160 | 6.6E-5 |
| NCBIfam | TIGR00839 | JCVI: aspartate ammonia-lyase | IPR004708 | Aspartate ammonia-lyase | 8 | 469 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.