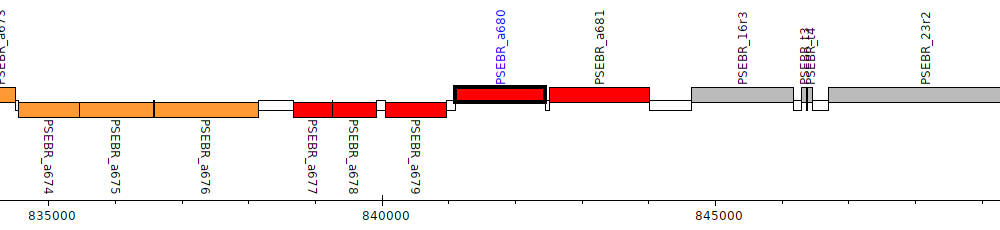
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a680
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas brassicacearum subsp. brassicacearum NFM421
GCF_000194805.1|latest |
| Locus Tag |
PSEBR_a680
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 841092 - 842441 (+ strand) |
Cross-References
| RefSeq | YP_004351832.1 |
| GI | 330807370 |
| Entrez | 10469533 |
| INSDC | AEA66828.1 |
| NCBI Locus Tag | PSEBR_a680 |
| UniParc | UPI0002041B2F |
| UniProtKB Acc | F2KBP6 |
| UniProtKB ID | F2KBP6_PSEBN |
| UniRef100 | UniRef100_A0A0D0RU85 |
| UniRef50 | UniRef50_Q9I700 |
| UniRef90 | UniRef90_P28269 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
beta-alanine--pyruvate transaminase
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -2.78 |
| Kyte-Doolittle Hydrophobicity Value | -0.084 |
| Molecular Weight (kDa) | 48.5 |
| Isoelectric Point (pI) | 6.63 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 3A8U | TRANSFERASE | 10/11/09 | Crystal Structure of omega-Amino Acid:Pyruvate Aminotransferase | Pseudomonas putida | 1.4 | X-RAY DIFFRACTION | 95.7 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 639 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000130 (850 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PSEBR_a680
Search term: beta-alanine--pyruvate transaminase
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
ornithine aminotransferase [Source:HGNC Symbol;Acc:HGNC:8091]
E-value:
7.0e-37
Percent Identity:
28.0
|
References
No references are associated with this feature.