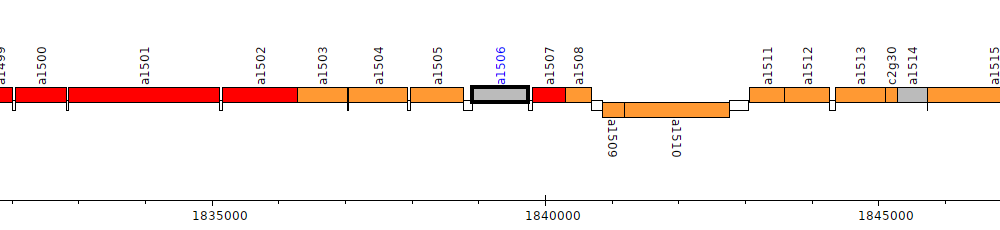
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a1506
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:SM00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pba02030 | Bacterial chemotaxis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01584 | CheW-like domain | IPR002545 | CheW-like domain | 126 | 258 | 6.0E-12 |
| SMART | SM00260 | chew_2 | IPR002545 | CheW-like domain | 122 | 257 | 6.3E-17 |
| PIRSF | PIRSF020479 | UCP020479_CheW | IPR014506 | Uncharacterised conserved protein UCP020479, signal transduction CheW-like | 1 | 272 | 1.1E-94 |
| SUPERFAMILY | SSF50341 | CheW-like | IPR036061 | CheW-like domain superfamily | 124 | 258 | 4.71E-15 |
| CDD | cd00588 | CheW_like | - | - | 128 | 256 | 2.54394E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.