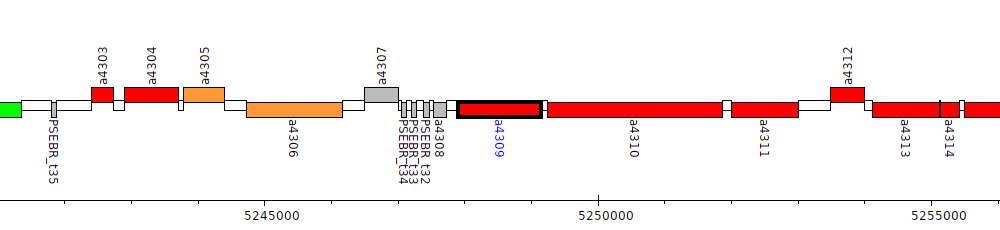
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a4309
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008652 | cellular amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00657
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009089 | lysine biosynthetic process via diaminopimelate |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000726
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004072 | aspartate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000726
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pba00270 | Cysteine and methionine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | spermidine biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-lysine biosynthesis VI | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pba00300 | Lysine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-lysine biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-methionine biosynthesis IV (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pba00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | grixazone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pba01210 | 2-Oxocarboxylic acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | norspermidine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pba01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba00261 | Monobactam biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 3-dehydroquinate biosynthesis II (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pba01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-lysine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00657 | JCVI: aspartate kinase | IPR001341 | Aspartate kinase | 65 | 404 | 5.2E-119 |
| SUPERFAMILY | SSF53633 | Carbamate kinase-like | IPR036393 | Acetylglutamate kinase-like superfamily | 4 | 243 | 4.06E-73 |
| Gene3D | G3DSA:3.30.2130.10 | - | - | - | 256 | 412 | 2.5E-69 |
| SUPERFAMILY | SSF55021 | ACT-like | IPR045865 | ACT-like domain | 250 | 328 | 2.84E-18 |
| PIRSF | PIRSF000726 | Asp_kin | IPR005260 | Aspartate kinase, monofunctional class | 58 | 411 | 0.0 |
| Gene3D | G3DSA:3.40.1160.10 | - | IPR036393 | Acetylglutamate kinase-like superfamily | 1 | 255 | 6.3E-108 |
| Pfam | PF01842 | ACT domain | IPR002912 | ACT domain | 269 | 326 | 1.9E-11 |
| FunFam | G3DSA:3.40.1160.10:FF:000002 | Aspartokinase | - | - | 1 | 255 | 2.1E-122 |
| FunFam | G3DSA:3.30.2130.10:FF:000002 | Aspartokinase | - | - | 254 | 412 | 1.1E-74 |
| CDD | cd04261 | AAK_AKii-LysC-BS | IPR041740 | Aspartokinase catalytic domain | 3 | 243 | 0.0 |
| SUPERFAMILY | SSF55021 | ACT-like | IPR045865 | ACT-like domain | 331 | 406 | 2.82E-24 |
| CDD | cd04923 | ACT_AK-LysC-DapG-like_2 | - | - | 343 | 405 | 1.47234E-30 |
| NCBIfam | TIGR00656 | JCVI: aspartate kinase, monofunctional class | IPR005260 | Aspartate kinase, monofunctional class | 1 | 405 | 0.0 |
| PIRSF | PIRSF000726 | Asp_kin | IPR005260 | Aspartate kinase, monofunctional class | 1 | 63 | 8.2E-23 |
| PANTHER | PTHR21499 | ASPARTATE KINASE | - | - | 1 | 403 | 1.7E-118 |
| CDD | cd04913 | ACT_AKii-LysC-BS-like_1 | - | - | 264 | 338 | 6.18519E-31 |
| Pfam | PF13840 | ACT domain | IPR027795 | CASTOR, ACT domain | 341 | 400 | 1.6E-14 |
| Pfam | PF00696 | Amino acid kinase family | IPR001048 | Aspartate/glutamate/uridylate kinase | 3 | 231 | 2.2E-49 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.