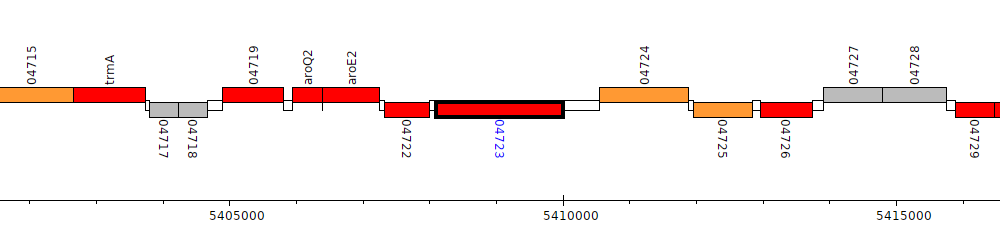
Pseudomonas sp. UW4, PputUW4_04723
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046565 | 3-dehydroshikimate dehydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02238
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0046279 | 3,4-dihydroxybenzoate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02238
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuu01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu00360 | Phenylalanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu00350 | Tyrosine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu00130 | Ubiquinone and other terpenoid-quinone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd08342 | HPPD_N_like | IPR041736 | 4-hydroxyphenylpyruvate dioxygenase, N-terminal | 297 | 425 | 1.23456E-27 |
| Coils | Coil | Coil | - | - | 64 | 91 | - |
| Pfam | PF14696 | Hydroxyphenylpyruvate dioxygenase, HPPD, N-terminal | - | - | 294 | 425 | 3.3E-32 |
| Gene3D | G3DSA:3.10.180.10 | - | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 435 | 630 | 8.4E-48 |
| Pfam | PF00903 | Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily | IPR004360 | Glyoxalase/fosfomycin resistance/dioxygenase domain | 438 | 560 | 5.8E-11 |
| CDD | cd07250 | HPPD_C_like | IPR041735 | 4-hydroxyphenylpyruvate dioxygenase, C-terminal | 436 | 622 | 6.55641E-66 |
| SUPERFAMILY | SSF54593 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 292 | 619 | 2.25E-63 |
| Pfam | PF01261 | Xylose isomerase-like TIM barrel | IPR013022 | Xylose isomerase-like, TIM barrel domain | 18 | 262 | 9.3E-43 |
| PANTHER | PTHR12110 | HYDROXYPYRUVATE ISOMERASE | - | - | 1 | 336 | 2.3E-95 |
| SUPERFAMILY | SSF51658 | Xylose isomerase-like | IPR036237 | Xylose isomerase-like superfamily | 1 | 269 | 7.32E-77 |
| Gene3D | G3DSA:3.20.20.150 | - | - | - | 1 | 269 | 8.3E-70 |
| Gene3D | G3DSA:3.10.180.10 | - | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 288 | 433 | 9.9E-40 |
| Hamap | MF_02238 | 3-dehydroshikimate dehydratase. | IPR043700 | 3-dehydroshikimate dehydratase | 1 | 625 | 46.12574 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.