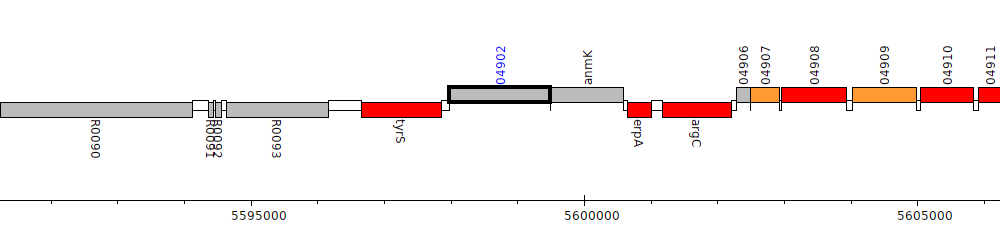
Pseudomonas sp. UW4, PputUW4_04902
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0042834 | peptidoglycan binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF04225
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:2.70.70.10:FF:000002 | Murein DD-endopeptidase MepM | - | - | 335 | 464 | 1.1E-50 |
| SUPERFAMILY | SSF51261 | Duplicated hybrid motif | IPR011055 | Duplicated hybrid motif | 209 | 459 | 5.75E-74 |
| Pfam | PF01551 | Peptidase family M23 | IPR016047 | Peptidase M23 | 362 | 458 | 4.3E-33 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 102 | 120 | - |
| Gene3D | G3DSA:3.10.450.350 | - | - | - | 133 | 221 | 2.3E-18 |
| Pfam | PF19425 | Csd3 second domain | IPR045834 | Csd3-like, second N-terminal domain | 229 | 349 | 5.8E-14 |
| Gene3D | G3DSA:2.70.70.10 | Glucose Permease (Domain IIA) | IPR011055 | Duplicated hybrid motif | 333 | 465 | 2.3E-91 |
| Gene3D | G3DSA:3.10.450.350 | - | - | - | 237 | 487 | 2.3E-91 |
| CDD | cd12797 | M23_peptidase | - | - | 363 | 448 | 8.95243E-42 |
| PANTHER | PTHR21666 | PEPTIDASE-RELATED | - | - | 125 | 461 | 7.5E-42 |
| Pfam | PF04225 | Opacity-associated protein A LysM-like domain | IPR007340 | Opacity-associated protein A | 138 | 215 | 1.6E-9 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 98 | 135 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.