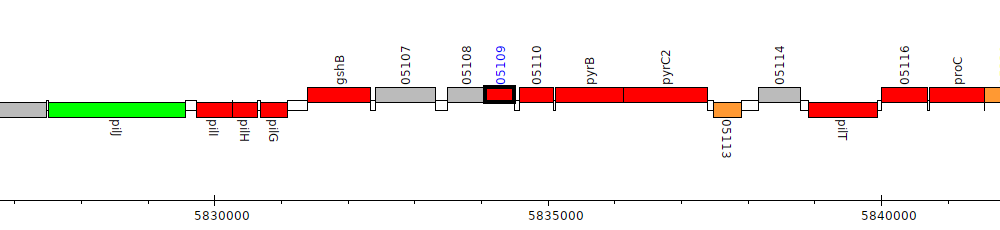
Pseudomonas sp. UW4, PputUW4_05109
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006364 | rRNA processing |
Inferred from Sequence Model
Term mapped from: InterPro:cd16964
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006139 | nucleobase-containing compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.420.140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53098 | Ribonuclease H-like | IPR012337 | Ribonuclease H-like superfamily | 4 | 138 | 6.98E-41 |
| Gene3D | G3DSA:3.30.420.140 | - | IPR037027 | YqgF/RNase H-like domain superfamily | 4 | 143 | 6.5E-42 |
| NCBIfam | TIGR00250 | JCVI: Holliday junction resolvase RuvX | IPR005227 | Putative pre-16S rRNA nuclease | 7 | 137 | 2.8E-43 |
| Hamap | MF_00651 | Putative pre-16S rRNA nuclease [yqgF]. | IPR005227 | Putative pre-16S rRNA nuclease | 4 | 140 | 24.691437 |
| SMART | SM00732 | rnase_8s | IPR006641 | YqgF/RNase H-like domain | 4 | 104 | 3.6E-41 |
| Pfam | PF03652 | Holliday junction resolvase | IPR005227 | Putative pre-16S rRNA nuclease | 5 | 138 | 1.1E-41 |
| CDD | cd16964 | YqgF | IPR005227 | Putative pre-16S rRNA nuclease | 6 | 138 | 7.4049E-59 |
| PANTHER | PTHR33317 | POLYNUCLEOTIDYL TRANSFERASE, RIBONUCLEASE H-LIKE SUPERFAMILY PROTEIN | IPR005227 | Putative pre-16S rRNA nuclease | 6 | 142 | 2.0E-24 |
| FunFam | G3DSA:3.30.420.140:FF:000002 | Putative pre-16S rRNA nuclease | - | - | 2 | 140 | 2.1E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.