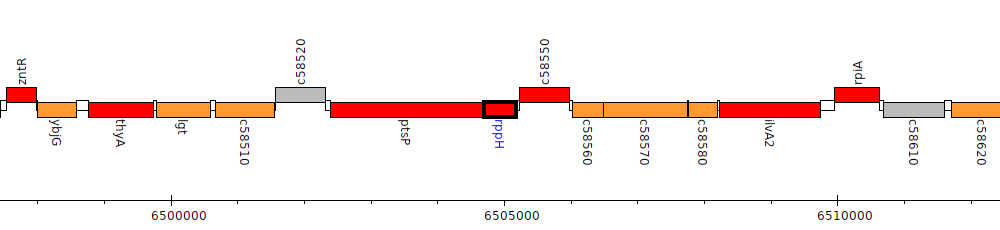
Pseudomonas protegens CHA0, PFLCHA0_c58540 (rppH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pprc03018 | RNA degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00298 | RNA pyrophosphohydrolase [rppH]. | IPR022927 | RNA pyrophosphohydrolase RppH | 1 | 156 | 37.547424 |
| PANTHER | PTHR43736 | ADP-RIBOSE PYROPHOSPHATASE | - | - | 2 | 151 | 5.8E-17 |
| Gene3D | G3DSA:3.90.79.10 | Nucleoside Triphosphate Pyrophosphohydrolase | - | - | 1 | 156 | 2.8E-46 |
| Pfam | PF00293 | NUDIX domain | IPR000086 | NUDIX hydrolase domain | 7 | 146 | 1.7E-26 |
| SUPERFAMILY | SSF55811 | Nudix | IPR015797 | NUDIX hydrolase-like domain superfamily | 4 | 155 | 2.3E-32 |
| PRINTS | PR00502 | NUDIX hydrolase family signature | IPR020476 | NUDIX hydrolase | 47 | 62 | 1.7E-6 |
| PRINTS | PR00502 | NUDIX hydrolase family signature | IPR020476 | NUDIX hydrolase | 33 | 47 | 1.7E-6 |
| CDD | cd03671 | Ap4A_hydrolase_plant_like | IPR022927 | RNA pyrophosphohydrolase RppH | 6 | 151 | 1.97431E-69 |
| FunFam | G3DSA:3.90.79.10:FF:000001 | RNA pyrophosphohydrolase | - | - | 1 | 159 | 3.0E-91 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.