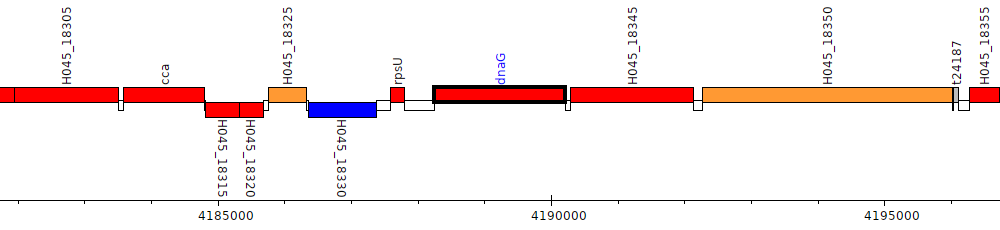
Pseudomonas poae RE_1-1-14, H045_18340 (dnaG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01807
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PF01807
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01807
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003896 | DNA primase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF08278
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006269 | DNA replication, synthesis of RNA primer |
Inferred from Sequence Model
Term mapped from: InterPro:PF08278
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016779 | nucleotidyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF10410
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppz03030 | DNA replication | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00766 | dnag_dnab_bind_seq8d | IPR013173 | DNA primase DnaG, DnaB-binding domain | 513 | 642 | 4.4E-31 |
| Pfam | PF08275 | DNA primase catalytic core, N-terminal domain | IPR013264 | DNA primase, catalytic core, N-terminal | 126 | 254 | 2.7E-44 |
| PANTHER | PTHR30313 | DNA PRIMASE | - | - | 1 | 647 | 0.0 |
| Gene3D | G3DSA:1.20.50.20 | DnaG, RNA polymerase domain, helical bundle | - | - | 371 | 427 | 7.0E-20 |
| FunFam | G3DSA:3.90.580.10:FF:000001 | DNA primase | - | - | 1 | 101 | 2.5E-42 |
| Pfam | PF13662 | Toprim domain | IPR006171 | TOPRIM domain | 261 | 327 | 2.3E-16 |
| CDD | cd03364 | TOPRIM_DnaG_primases | IPR034151 | Bacterial DnaG primase, TOPRIM domain | 261 | 342 | 2.30695E-34 |
| Pfam | PF01807 | CHC2 zinc finger | IPR002694 | Zinc finger, CHC2-type | 6 | 100 | 2.5E-41 |
| FunFam | G3DSA:3.40.1360.10:FF:000002 | DNA primase | - | - | 243 | 370 | 4.8E-53 |
| SUPERFAMILY | SSF57783 | Zinc beta-ribbon | - | - | 4 | 101 | 6.12E-38 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 478 | 497 | - |
| FunFam | G3DSA:3.90.980.10:FF:000001 | DNA primase | - | - | 110 | 242 | 1.1E-49 |
| SUPERFAMILY | SSF56731 | DNA primase core | - | - | 116 | 427 | 3.66E-109 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 467 | 497 | - |
| Gene3D | G3DSA:3.90.980.10 | - | IPR037068 | DNA primase, catalytic core, N-terminal domain superfamily | 113 | 242 | 2.1E-49 |
| Gene3D | G3DSA:3.40.1360.10 | - | - | - | 243 | 370 | 2.9E-41 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 94 | 114 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 94 | 108 | - |
| Pfam | PF10410 | DnaB-helicase binding domain of primase | IPR019475 | DNA primase, DnaB-helicase binding domain | 373 | 425 | 1.4E-8 |
| NCBIfam | TIGR01391 | JCVI: DNA primase | IPR006295 | DNA primase, DnaG | 5 | 419 | 0.0 |
| Gene3D | G3DSA:1.10.860.10 | DNAb Helicase; Chain A | IPR016136 | DNA helicase DnaB, N-terminal/DNA primase DnaG, C-terminal | 504 | 651 | 3.4E-33 |
| SMART | SM00493 | toprim5 | IPR006171 | TOPRIM domain | 261 | 332 | 1.0E-19 |
| SMART | SM00400 | primzinc3 | IPR002694 | Zinc finger, CHC2-type | 36 | 90 | 1.7E-33 |
| Gene3D | G3DSA:3.90.580.10 | - | IPR036977 | DNA Primase, CHC2-type zinc finger | 2 | 102 | 2.8E-36 |
| SUPERFAMILY | SSF117023 | DNA primase DnaG, C-terminal domain | - | - | 513 | 639 | 7.98E-31 |
| PIRSF | PIRSF002811 | DnaG | IPR030846 | DNA primase DnaG, bacteria | 2 | 648 | 0.0 |
| Pfam | PF08278 | DNA primase DnaG DnaB-binding | IPR013173 | DNA primase DnaG, DnaB-binding domain | 517 | 633 | 1.2E-24 |
| Hamap | MF_00974 | DNA primase [dnaG]. | IPR030846 | DNA primase DnaG, bacteria | 4 | 643 | 30.002378 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.