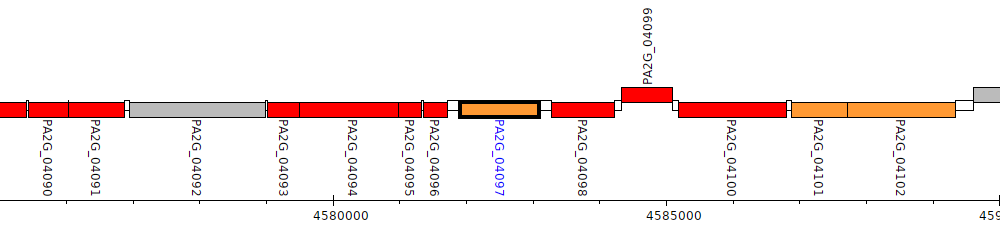
Pseudomonas aeruginosa 2192, PA2G_04097
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004888 | transmembrane signaling receptor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF58104 | Methyl-accepting chemotaxis protein (MCP) signaling domain | - | - | 88 | 334 | 3.53E-37 |
| SMART | SM00283 | MA_2 | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 95 | 351 | 3.5E-5 |
| Gene3D | G3DSA:1.10.287.950 | - | - | - | 60 | 339 | 8.7E-42 |
| PANTHER | PTHR32089 | METHYL-ACCEPTING CHEMOTAXIS PROTEIN MCPB | - | - | 85 | 314 | 3.4E-33 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 195 | 222 | 1.4E-16 |
| Pfam | PF00015 | Methyl-accepting chemotaxis protein (MCP) signalling domain | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 169 | 315 | 4.1E-28 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 224 | 253 | 1.4E-16 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.