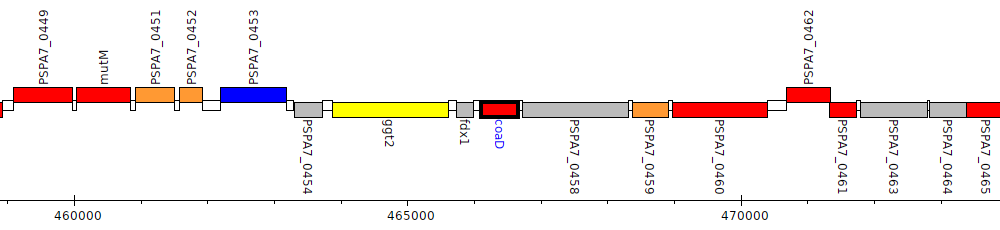
Pseudomonas aeruginosa PA7, PSPA7_0457 (coaD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008771 | [citrate (pro-3S)-lyase] ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00764
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004595 | pantetheine-phosphate adenylyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd02163
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015937 | coenzyme A biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd02163
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00125
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00125
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pap01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap00770 | Pantothenate and CoA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | coenzyme A biosynthesis II (eukaryotic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01020 | Lipopolysaccharide core biosynthesis protein signature | IPR001980 | Phosphopantetheine adenylyltransferase | 44 | 65 | 4.2E-50 |
| NCBIfam | TIGR00125 | JCVI: cytidyltransferase-like domain | IPR004821 | Cytidyltransferase-like domain | 27 | 87 | 1.9E-19 |
| PRINTS | PR01020 | Lipopolysaccharide core biosynthesis protein signature | IPR001980 | Phosphopantetheine adenylyltransferase | 73 | 97 | 4.2E-50 |
| PRINTS | PR01020 | Lipopolysaccharide core biosynthesis protein signature | IPR001980 | Phosphopantetheine adenylyltransferase | 26 | 44 | 4.2E-50 |
| Hamap | MF_00151 | Phosphopantetheine adenylyltransferase [coaD]. | IPR001980 | Phosphopantetheine adenylyltransferase | 26 | 182 | 36.503994 |
| SUPERFAMILY | SSF52374 | Nucleotidylyl transferase | - | - | 25 | 180 | 7.24E-47 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 20 | 183 | 4.7E-62 |
| CDD | cd02163 | PPAT | IPR001980 | Phosphopantetheine adenylyltransferase | 27 | 179 | 4.89004E-100 |
| NCBIfam | TIGR01510 | JCVI: pantetheine-phosphate adenylyltransferase | IPR001980 | Phosphopantetheine adenylyltransferase | 28 | 180 | 6.1E-59 |
| Pfam | PF01467 | Cytidylyltransferase-like | IPR004821 | Cytidyltransferase-like domain | 29 | 157 | 1.3E-23 |
| PANTHER | PTHR21342 | PHOSPHOPANTETHEINE ADENYLYLTRANSFERASE | - | - | 26 | 182 | 4.6E-45 |
| PRINTS | PR01020 | Lipopolysaccharide core biosynthesis protein signature | IPR001980 | Phosphopantetheine adenylyltransferase | 136 | 158 | 4.2E-50 |
| SMART | SM00764 | citrate_ly_lig5 | IPR013166 | Citrate lyase ligase, C-terminal | 36 | 177 | 0.0096 |
| PRINTS | PR01020 | Lipopolysaccharide core biosynthesis protein signature | IPR001980 | Phosphopantetheine adenylyltransferase | 109 | 125 | 4.2E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.